Causal Networks
As we discussed in the last post, Bayesian networks can often be written in multiple equivalent ways by reversing some of the arrows. Generally the most natural choice has arrows matching the causal direction of the effects. A Bayesian network in which the arrows have causal meaning is a causal network. In this post we’ll build up to a clearer idea of what that means.
Consider again the models A and B of IQ(X) vs log-income(Y) from the last post. They’re GBNs, so we can write them as systems of linear models as follows.
\[ \begin{align*} X &= \beta_{10} + \epsilon_X \\ Y &= \beta_{20} + \beta_{21}X + \epsilon_Y \end{align*} \]
\[ \begin{align*} X &= \beta_{10} + \beta_{11}Y + \epsilon_X \\ Y &= \beta_{20} + \epsilon_Y \end{align*} \]
Where \(\epsilon_X\) and \(\epsilon_Y\) are normally distributed error terms that represent any deviation between the linear models and the observed data, as in linear regression.
This form of models A and B is called a structural equation model (SEM). These are given a causal interpretation, where the variables on the right hand side (dependent variables) of an equation cause the variable on the left (independent variable). This causal information is not preserved with algebraic maniupulation, e.g. substituting the equation for X into Y’s equation in model A isn’t valid.
In the last post we had data generated by an opaque process and we fit our model parameters to the data. Now we’ll take another view by using our models to generate data. We can do this in R by creating a pipeline of random samples. For model A, for example, we’ll draw from X first and each of these draws lets us compute the correct mean for the normal distribution to draw Y from. We’ll call this sort of pipeline a data generating process. There’s nothing special about code, cause and effect processes in nature can also be thought of as data generating processes.
Let’s look at that code. We’re using the parameter values we learned from data in our last post.
# Set a RNG seed, so our simulations will give the same results every time we run our code
set.seed(2413)
# The number of samples we'll use for our simulations.
n <- 1000
#
eps_X.A <-rnorm(n, 0, 15) # Draw n random samples from a normal distribution with mean=0, standard deviation=15
eps_Y.A <- rnorm(n, 0, 1)
X.A <- 100 + eps_X.A
Y.A <- 4 + X.A/60 + eps_Y.A
#
eps_X.B <- rnorm(n, 0, sqrt(60^2/17))
eps_Y.B <- rnorm(n, 0, sqrt(17/16))
Y.B <- 17/3 + eps_Y.B
X.B <- 80 + 60*Y.B/17 + eps_X.B
# simulations can also be done from a fitten BN, see ?rbn(X.A, Y.A) and (X.B, Y.B) have the same joint distribution, and the same distribution as the data generated directly from a MVN in the last post. Therefore, any predictions made based on observations will be the same (e.g. \(P_A(X|Y)=P_B(X|Y)\)). When the arrows are given causal meaning, the models differ for predictions about interventions.
An intervention is the smallest possible change to a causal network that forces a variable to take a specific value. For a network structure, this means removing all arrows pointing at the intervened variable. For SEM, the target equation is replaced with a constant. For our simulation code, a random draw is replaced with a constant.
Interventions are written as e.g. \(\mathrm{do(X = x)}\) The probability distribution for a variable given that another variable was intervened on is written e.g. \(P(Y\,|\,\mathrm{do}(X = x))\). Other equivalent notations are \(P(Y\,|\,\hat{x})\) and \(P(Y_x)\), though we won’t use these.
Interventions generally change the distribution of a network. This change is not in general the same among observationally equivalent networks. e.g. in the case of our models A and B, \(P_A(Y\,|\,\mathrm{do}(X = x))\neq P_B(Y\,|\,\mathrm{do}(X = x))\)
This is what it looks like if we intervene with \(\mathrm{do}(Y = 20)\) (e.g. an annual income of \(\$10^{20}\)(!)), for each of the three representations of a causal network we covered.
Network Structure
library(bnlearn)mdl.A <- model2network('[X][Y|X]')
mdl.B <- model2network('[X|Y][Y]')
# 'mutilated' is for interventions, it sets nodes to constants and cuts their incoming arrows
mut.A <- mutilated(mdl.A, list(Y = 20))
mut.B <- mutilated(mdl.B, list(Y = 20))
mut.B$arcs <- rbind(mut.B$arcs) # 'mutilated' is broken when returning one edge, need to cast edges to matrixpar(mfrow=c(2, 2))
plot(mdl.A, main='A')
plot(mut.A, main='A | do(Y = 20)')
plot(mdl.B, main='B')
plot(mut.B, main='B | do(Y = 20)')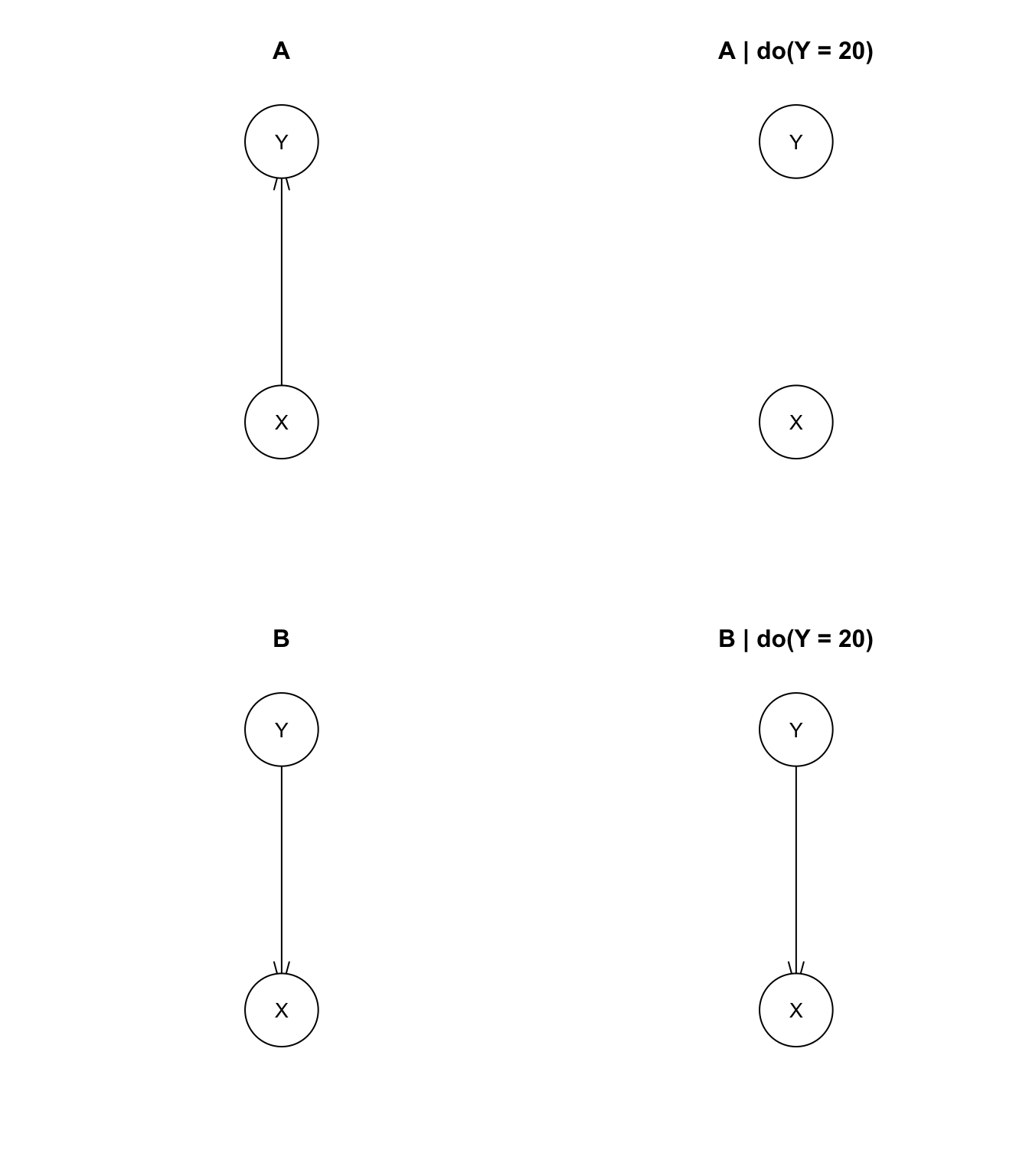
SEM
(A | do(Y = 20)) \[ \begin{align*} X &= \beta_{10} + \epsilon_X \\ Y &= 20 \end{align*} \]
(B | do(Y = 20)) \[ \begin{align*} X &= \beta_{10} + \beta_{11}Y + \epsilon_X \\ Y &= 20 \end{align*} \]
Data generating process
set.seed(544)
#
eps_X.A.mut <-rnorm(n, 0, 15)
eps_Y.A.mut <- rnorm(n, 0, 1)
X.A.mut <- 100 + eps_X.A.mut
Y.A.mut <- rep(20, n) # Intervention
#
eps_X.B.mut <- rnorm(n, 0, sqrt(60^2/17))
eps_Y.B.mut <- rnorm(n, 0, sqrt(17/16))
Y.B.mut <- rep(20, n) # Intervention
X.B.mut <- 80 + 60*Y.B.mut/17 + eps_X.B.mutmean(X.A.mut)## [1] 99.26795mean(X.B.mut)## [1] 150.1895Expected IQ stays the same under model A, but increases by 50 points under model B. Model A more closely matches what we’d expect in reality.
We’ve kept things simple by sticking to two variables. In the next post we’ll examine causal interactions between three variables.